Translate this page into:
Immobilization of Urease on DEAE-Cellulose Strips for One Step Urea Detection
Address for correspondence Arvind M. Kayastha, FNASc, School of Biotechnology, Institute of Science, Banaras Hindu University, Varanasi 221005, Uttar Pradesh, India (e-mail: kayasthabhu@gmail.com).
This article was originally published by Thieme Medical and Scientific Publishers Pvt. Ltd. and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Urease isolated from pigeonpeas was immobilized on 1 cm × 6 cm diethylaminoethyl (DEAE)-cellulose strips. The optimum immobilization (55% activity) was observed at 4°C, with a protein concentration of 1 mg per strip at pH 6.5. Immobilized strips stored at 4°C showed an increased shelf-life and there was no leaching of enzyme from the immobilized strips for 6 months. Enzyme could be eluted on application of 0.2 M KCl in buffer, which showed that the binding of the enzyme was tight on DEAE-cellulose strips. These strips when used along with dye, phenol red (coimmobilized) gave different shades from light yellow to dark magenta depending upon the concentration of urea. The ease of immobilization of urease on DEAE-cellulose strips and easy availability of pigeonpeas urease described in the present study makes it a suitable product for potential applications in urea diagnostics.
Keywords
diethylaminoethyl-cellulose paper strips
urease
urea
immobilization
analytical application
Introduction
Estimation of urease is frequently performed in the medical as the metabolic function of the kidney is reflected in the concentration of urea in blood or urine.1 Urease (urea amidohydrolase, EC 3.5.1.5) catalyzes the hydrolysis of urea yielding ammonia and carbon dioxide.2,3 It has now become possible to decompose urea using artificial kidneys containing immobilized urease (Japanese Patent Publications numbers 36751/85 and 17467/86). In food industry, urea is undesirable, particularly in biologically fermented food products, such as beer, wine, soy sauce, or sake. For these applications, neutral urease obtained from seeds of plant bean species such as canavalia ensiformis, or from microorganisms such as eubacterium aerofaciens and P. mirabilis are very useful and yielding good results.4 There is ongoing approach to explore cheap sources of urease, which are readily available and have better physico-chemical properties.
The utilization of any enzyme is driven by its versatility, regio-, chemo- and enantio-selectivity while operating under mild conditions making any process environmentally compatible.5 Success of an enzyme's utilization in industries depends on its reuse. Enzyme immobilization is one of the best ways to reuse the enzymes. The anchoring of an enzyme to a solid insoluble support should be straightforward and cost-efficient. The key factor determining the success or failure of immobilization depends on the methodology used for enzyme immobilization.6 One of the serious problems for the industrial application of the technology of immobilization is the diffusional limitations to the transport of substrate and product.7 This limitation arises from the stearic hindrance of the solid matrix to the free diffusion of substrates and products toward or away from the catalytic site of the immobilized enzymes.
Urease from pigeonpea was purified and characterized from our laboratory8 and can be bound on DEAE-cellulose strips at neutral pH. The ready availability of urease, its low cost, and the ease of its immobilization on cellulose paper strips described in the present study, makes it a product for future applications in therapeutics and diagnostics.
Materials and Methods
Materials
Urease was isolated from dehusked pigeonpeas procured from agriculture shops from the local market. Urea (enzyme grade), ammonium chloride, and Tris were obtained from Sisco Research Laboratoties (Mumbai, India). Nessler's reagent and phenol red were obtained from HiMedia Laboratories (Mumbai, India). Tricholoacetic acid was obtained as 20% solution from Loba Chemie (India). DEAE-cellulose paper (DE 81) from Whatman International (Maidstone, Kent, United Kingdom) was a generous gift from Professor PM Dey, School of Biological Sciences, University of London, United Kingdom. All solutions were prepared in Milli Q water (Millipore, United States).
Urease Purification
Urease was extracted in 0.1 M Tris-acetate buffer, pH = 6.8 and purified 200 fold to electrophoretic homogeneity, from dehusked pigeonpeas according to the method described from this laboratory.8 The specific activity of the purified enzyme was 5 × 103 units/mg of protein, with approximately 10% yield.
Protein Assay
Protein content was measured using the method of Lowry et al,9 using bovine serum albumin as standard protein.
Urease Assay
For measurement of urease activity, ammonia liberated on incubating the enzyme with urease in a fixed time period at an enzyme saturating concentration of urease was determined using Nessler's reagent. The golden brown color produced was measured spectrophotometrically at 405 nm. The amount of ammonia liberated in the test solution was calculated by calibrating the reagent with standard ammonium chloride solution. An enzyme unit was defined as the amount of enzyme required to liberate 1 μmole of ammonia per minute under test conditions (0.1 M urea, 0.05 M Tris-acetate buffer, pH = 7.3 at 37°C).
Immobilization on Paper Strips and Assay
The DEAE-cellulose paper was cut into 1 cm × 6 cm strips and prewashed with Tris-acetate buffer (0.1 M, pH = 6.5) and dried using a hair drier. Strips were dipped in a solution of urease 1 mg/mL and 0.6 mg/mL phenol red and dried again and stored in capped tubes till used at 4°C. By doing so, urease and dye were coimmobilized on the DEAE-cellulose strips. These strips were then dipped in different concentrations of urea (0.1–2 g/L). This yields different colors on strips from light yellow to deep magenta. Response time for this was only 30 seconds. Such a strip chart developed can be then used for assay of urea in a blood serum samples.
Storage Stability
DEAE-cellulose strips (1 cm × 6 cm) were stored at 4°C for several months and used, when required.
Results and Discussion
The immobilization of urease on DEAE-cellulose strips under different conditions of immobilization was studied. The results are obtained on the percentage immobilization with varying protein concentration and optimal volume loading were studied and are summarized in ►Table 1. clearly showed that optimum immobilization was obtained at 1 mg protein per strip at 4°C. Immobilization percentage reduced above or below 1 mg of protein/strip. Optimal loading of volume was 80 μL and the optimum incubation time was 24 hours. A pH of 6.5 was maintained for immobilization so that urease is tightly bound to DEAE-cellulose. The shelf-life of immobilized strips at 4°C increased to about 6 months (data not shown). It may be noted here that during urease purification from pigeonpeas, a DEAE-cellulose chromatography was an essential step and the urease adsorbed could be eluted only by applying a KCl gradient, which confirmed that urease is tightly bound to DEAE-cellulose.8
| Condition | Volume of urease loaded (μL) | Protein conc. (mg/mL) | Incubation (h) | Immobilization (%) |
|---|---|---|---|---|
| Protein concentration (mg/mL) | ||||
| 0.1 | 80 | – | 24 | 45 |
| 0.2 | 80 | – | 24 | 48 |
| 0.5 | 80 | – | 24 | 50 |
| 1.0 | 80 | – | 24 | 55 |
| 1.5 | 80 | – | 24 | 52 |
| Volume of enzyme loaded (μL) | ||||
| 10 | – | 1 | 24 | 40 |
| 20 | – | 1 | 24 | 45 |
| 40 | – | 1 | 24 | 50 |
| 80 | – | 1 | 24 | 55 |
| 100 | – | 1 | 24 | 53 |
| Incubation (h) | ||||
| 6 | 80 | 1 | – | 45 |
| 12 | 80 | 1 | – | 50 |
| 24 | 80 | 1 | – | 55 |
| 48 | 80 | 1 | – | 45 |
| 60 | 80 | 1 | – | 38 |
Abbreviation: DEAE, diethylaminoethyl.
Highly purified urease was successfully immobilized on DEAE-cellulose paper strips. The percentage retention was 55% and the binding was tight. This was confirmed by elution of the enzyme by using 0.2 M KCl in buffer. No enzyme was eluted when strips were washed with buffer alone. The levels of urea in the clinical range are easily detectable by this method. The urease strips are highly specific for detection of urea present in serum. Earlier studies have clearly showed that urease has an absolute specificity for its substrate urea.8 Strips give different shades of color depending upon the concentration of urea present in the serum samples. It has a color range from light yellow to dark magenta. Normal values of urea in blood/serum are 0.15 to 0.40 g/L and the dynamic range is 0.5 to 2 g/L. The color chart of urea strip was prepared and is shown in ►Fig. 1. The control strip showed pale yellow color. Though many different methods are available for urea assay, present method provides a simple and fast detection based on the color matching with the strip.
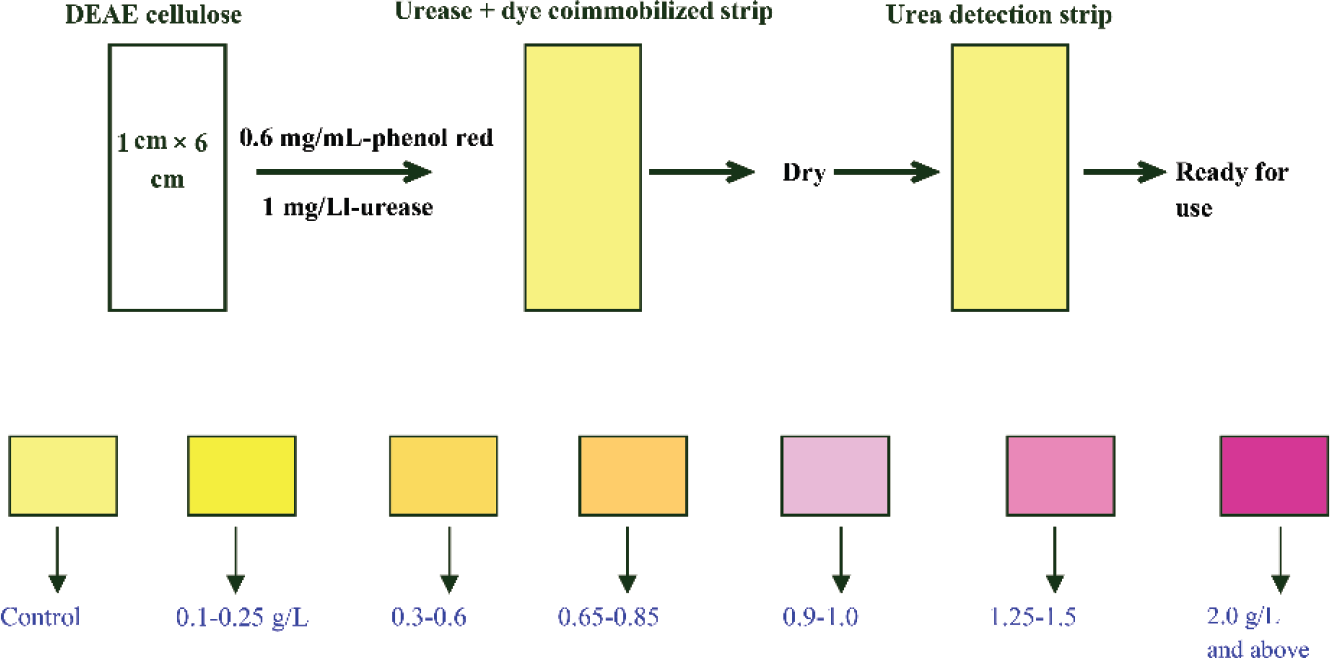
- The one step urea detection: the top panel shows the preparation of the urease strip and the bottom panel shows the color chart upon addition of different concentrations of urea.
Immobilization led to stabilization of urease on the strip with a shelf-life of 6 months at 4°C. Similar storage stability was observed earlier when urease was immobilized on gold nanoparticles; 4 however, cost-wise DEAE-cellulose paper strips are much less expensive. We also immobilized urease on glutaraldehyde-activated chitosan beads; however, getting uniform sized chitosan beads was a problem and the shelf-life was approximately 1 month.10 Therefore, these cellulose strips may have potential market for urea assay. Furthermore, detection of urea in milk samples is important to detect ‘adulteration in milk’ in several parts of western Uttar Pradesh. These strips may be useful for detection of urea in ‘synthetic milk’ and thus to check its adulteration.
Acknowledgments
A.M.K. is thankful to Dr. K. Ravi Charan Reddy for help with enzyme preparation from dehusked pigeonpeas and Prof. P. M Dey, School of Biological Sciences, University of London, United Kingdom for kindly providing the DEAE-cellulose paper from Whatman's United Kingdom. Laboratory facility from School of Biotechnology, Banaras Hindu University, is greatly acknowledged.
Conflict of Interest
None declared.
References
- Immobilization of urease from pigeonpea (Cajanus cajan L .) in polyacrylamide gels and calcium alginate beads. Biotechnol Appl Biochem. 1998;27(01):25-29.
- [CrossRef] [PubMed] [Google Scholar]
- Molecular biology of microbial ureases. Microbiol Rev. 1995;59(03):451-480.
- [CrossRef] [PubMed] [Google Scholar]
- Response surface analysis of nano-ureases from canavalia ensiformis and cajanus cajan. Int J Biol Macromol. 2011;49(04):674-680.
- [CrossRef] [PubMed] [Google Scholar]
- Understanding enzyme immobilisation. Chem Soc Rev. 2009;38(02):453-468.
- [CrossRef] [PubMed] [Google Scholar]
- Comparative study of solid silica nanoparticles and hollow silica nanoparticles for immobilization of lysozyme. Chem Eng J. 2008;137:38-44.
- [CrossRef] [Google Scholar]
- Novel synthesis of ferric impregnated silica nanoparticles and their evaluation as a matrix for enzyme immobilization. J Phys Chem. 2007;111:3842-3847.
- [CrossRef] [Google Scholar]
- Optimal immobilization of β-galactosidase from Pea (PsBGAL) onto Sephadex and chitosan beads using response surface methodology and its applications. Bioresour Technol. 2009;100(10):2667-2675.
- [CrossRef] [PubMed] [Google Scholar]
- Purification and characterization of urease from dehusked pigeonpea (Cajanus cajan L) seeds. Phytochemistry. 2002;61(05):513-521.
- [CrossRef] [PubMed] [Google Scholar]
- Protein measurement with the folin phenol reagent. J Biol Chem. 1951;193(01):265-275.
- [CrossRef] [PubMed] [Google Scholar]
- Pigeonpea (Cajanus cajan L .) urease immobilized on glutaraldehyde-activated chitosan beads and its analytical applications. Appl Biochem Biotechnol. 2001;96((01) (03)):41-53.
- [CrossRef] [PubMed] [Google Scholar]





